BrainJ
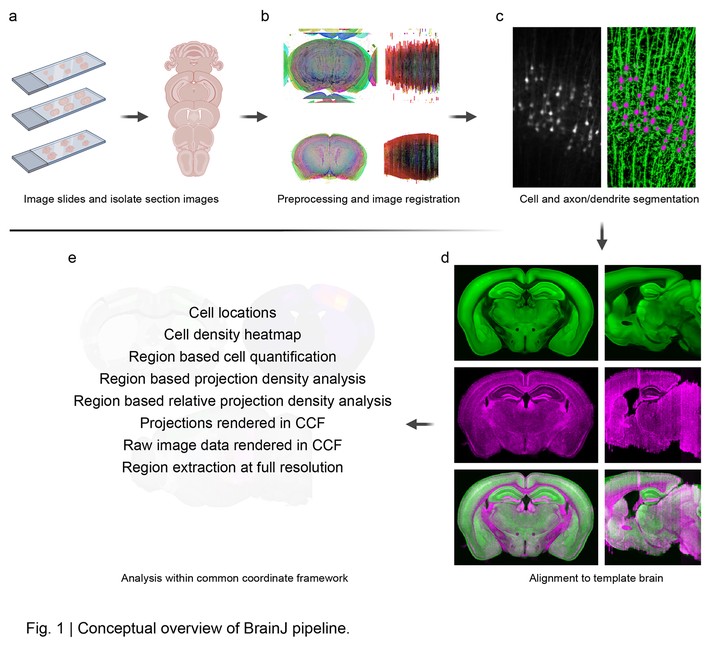
BrainJ enables high-throughput analysis of serial tissue sections imaged using confocal or widefield techniques. Developed in Fiji, our approach leverages freely available tools for machine learning pixel classification for cell detection and mesoscale mapping of axons and dendrites. With a simple graphical user interface, this approach is easy to deploy and use, requiring no coding experience and minimal manual intervention to process multiple datasets. Our approach is extensible to any project requiring serial section reconstruction and atlas-based analysis and allows a typical whole brain dataset to be reconstructed and analyzed within ~4 hours.
Project Author(s)
Luke Hammond
Project Links
https://github.com/lahammond/BrainJ
Project Video
This post was automatically generated by Luke Hammond